multiplex wormhole
Explore Optimization Parameters
The optimization process relies on a simulated annealing algorithm which allows earlier iterations to accept increases in dimer load to overcome local optima. The probability of accepting increased dimer load is defined by a variety of parameters, which can be explored with this function.
See here for optimization process details included simulated annealing algorithm parameterization.
Usage
The plot_ASA_temps.py function is setup to accept two different sets of inputs: 1) Provided simulated annealing parameters, or 2) Provided datasets used to calculate simulated annealing parameters (the default used in optimize_multiplex.py). Inputs are shown for both options. Provided datasets are ignored if MAX_DIMER and MIN_DIMER are provided.
Python syntax
import multiplex_wormhole as mw
# provided parameters (with defaults shown):
mw.plotASAtemps(OUTPATH, MIN_DIMER, MAX_DIMER, DECAY_RATE=0.95, T_INIT=None, T_FINAL=0.1, DIMER_ADJ=0.1, PROB_ADJ=2):
# provided datasets using default parameters:
mw.plotASAtemps(OUTPATH, PRIMER_FASTA, DIMER_SUMS, DIMER_TABLE, N_LOCI, KEEPLIST=None, SEED=None, BURNIN=100, deltaG=False)
# providing a dataset with user-defined paramers:
mw.plotASAtemps(OUTPATH, PRIMER_FASTA, DIMER_SUMS, DIMER_TABLE, N_LOCI, KEEPLIST=None, SEED=None, BURNIN=100,
deltaG=False, DECAY_RATE=0.95, T_INIT=None, T_FINAL=0.1,PROB_ADJ=2)
# providing a previous set of primers (e.g., from previous optimization) rather than new data, with all other defaults:
mw.plotASAtemps(OUTPATH, PRIMER_FASTA, SEED, DIMER_SUMS, DIMER_TABLE, N_LOCI, KEEPLIST=None)
Command line syntax
cd ~/multiplex_wormhole/src/multiplex_wormhole #move into directory holding scripts
# provided parameters:
python3 plot_ASA_temps.py -o OUTPATH [-m MIN_DIMERS] [-a MAX_DIMERS] [-r DECAY_RATE] [-i TEMP_INIT] [-f TEMP_FINAL] [-p PROB_ADJ] [-g]
# providing datasets:
python3 plot_ASA_temps.py -o OUTPATH [-f PRIMER_FASTA] [-d DIMER_SUMS] [-t DIMER_TABLE] [-n NLOCI] [-k KEEPLIST] [-b BURNIN] [-z SEED] [-g]
Arguments
OUTPATH (-o) : Prefix for output files, including directory path.
PRIMER_FASTA (-f) : FASTA filepath containing primer sequences to test. Important: PrimerIDs in this file must match primer pair IDs in DIMER_SUMS and DIMER_TABLE! (Default: None)
DIMER_SUMS (-d) : Filepath to CSV containing dimer loads per primer pair. (Default: None)
DIMER_TABLE (-t) : Filepath to CSV containing pairwise primer dimer loads. (Default: None)
N_LOCI (-n) : Target panel size (includes keeplist loci), i.e. the number of unique targets and primer pairs in the optimized multiplex. (Default: None)
KEEPLIST (-k) : Filepath to FASTA file containing primer sequences that are required to be included in final primer set. (Default: None)
SEED (-e) : CSV to primer set used as initial loci, in format of multiplex_wormhole output *_primers.csv. This option overrides N_LOCI, so the number of loci in the SEED set will be the final number of loci in the primer set. (Default: None)
MIN_DIMER (-m) : Minimum ‘bad’ dimer change expected going from one iteration to the next. Generally MIN_DIMER=1. (Default: None - calculated from data)
MAX_DIMER (-a) : Maximum ‘bad’ dimer change expected moving from one iteration to the next. (Default: None - calculate from data)
DECAY_RATE (-r) : Base value used in decay function for temperature. Values close to 1 lead to slow temperature decay while decreasing values lead to more rapid temperature decay. (Default: 0.95, limit: 0-1)
T_INIT (-i) : Initial temperature where simulated annealing starts. If not provided, this value will be calculated from changes in dimer load. (Default: None- set adaptively)
Adaptively calculated as MIN_DIMER + DIMER_ADJ * (MAX_DIMER - MIN_DIMER)
T_FINAL (-j) : Final temperature where simulated annealing ends. If not provided, this value will be calculated from changes in dimer load. (Default: 0.1)
BURNIN (-b) : Number of ‘bad’ swaps used to sample changes in dimer loads. (Default: 100)
PROB_ADJ (-p) : Multiplier to adjust dimer acceptance probability, where 1=no adjustment and higher values reduce dimer acceptance probability. (Default: 2)
The probability of accepting an increase in dimer load follows e^(-PROB_ADJ*cost / temperature)
deltaG (-g) : Optimize for total dimer tally (False) or maximum average deltaG among dimers (True)? [Default: False]
Outputs
Probabilities reflect default parameters, although the initial (higher) temperature will vary depending on the dataset.
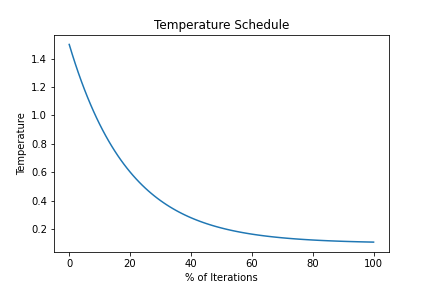
OUTPATH_TemperatureSchedule.png
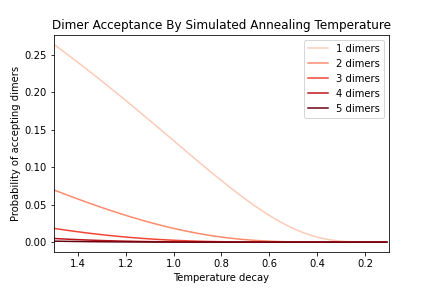
OUTPATH_DimerAcceptanceByTemp.png
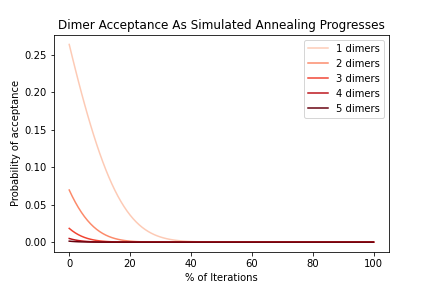
OUTPATH_DimerAcceptanceByIteration.png